The projects described below are featured to:
(a) highlight the variety of research opportunities available in the CSU Chemistry program, and
(b) link less-experienced undergraduate researchers to thoughtfully constructed assignments, with the aim to maximize discoveries and research productivity.
Please note: prospective REU participants are welcome to apply to work with any of our participating faculty.
Mentors: Professors Jeff Bandar and Eugene Chen
Project Description:
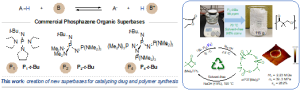
Organic superbases are neutral compounds that possess very strong basicity and display unique properties compared to traditional anionic bases. The Bandar and Chen Groups at CSU employ these bases as catalysts for the synthesis of bioactive small molecules and recyclable polymers with closed-loop chemical circularity, respectively. These protocols rely on commercially available superbases, but to improve this chemistry, the invention of new superbases is critical. Students involved in this project will be mentored by senior graduate students in the Bandar and Chen labs to accomplish this task. The research will involve the synthesis of new superbases, analysis of their basicity and optimization of their catalytic performance for the sustainable synthesis of pharmaceuticals and circular polymers.
Description of REU Student Activities:
- This student will work with their mentors to design and prepare new superbases, then test and optimize their performance for catalyzing the synthesis of small molecules and polymers
- Students will conduct daily laboratory work, including learning proper and safe techniques for reaction setup, reaction analysis, compound purification and the use of all associated instruments and equipment
- Individual mentorship from senior graduate students on planning, conducting and analyzing organic synthesis experiments
- Participation in and presentations at Bandar and Chen group meetings
- Practice activities and preparation for public scientific presentations and writing
Mentors: Professors Eugene Chen and Garret Miyake
Project Description:
Synthetic polymers are among the most important materials to modern society. However, current plastics aren’t successfully disposed of or recycled presenting a tremendous concern for the environment and human health as well as a waste of valuable resources. To address this challenge, this project focuses on the design, discovery, and development of next-generation circular polymers that are designed for chemical recyclability. These plastics have the potential to replace today’s non-recyclable or difficult-to-recycle plastics.
Description of REU Student Activities:
- Synthesis of chemically recyclable plastics
- Characterization of materials properties of plastics
- Development of conditions for chemical recycling
- Participate in Chen and Miyake group meetings

Mentors: Professors Seonah Kim and Garett Miyake
Project Description:
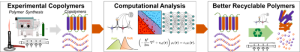
Plastic waste is an urgent issue globally, but the diverse applications of modern plastics make addressing this waste challenging. Copolymers, in particular, present a challenging recycling problem due to their complex chemical and morphological properties, with few tools to predict copolymer properties in silico. In this project, we will explore copolymer properties through computations and experiments to improve recyclable copolymers. The REU student will gain valuable experience in experimental polymer chemistry, using standard air-free methods and polymerization methods such as ROMP (Ring-Opening Metathesis Polymerization) to synthesize and characterize a wide variety of recyclable copolymers. The REU student will also use cutting-edge computational methods such as density functional theory (DFT) and machine learning (ML) to model real-world copolymers, focusing on property prediction via ML.
Description of REU Student Activities:
- Building polymer property database and developing machine learning models
- Recyclable polymer design and synthesis
- Python programming and data science
- Experimental polymer property measurements and analysis

Mentors: Professors Seonah Kim and Robert Paton
Project Description:
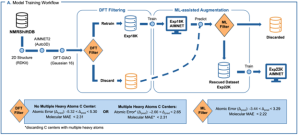
This project introduces a machine learning (ML) methodology to improve the prediction of carbon-13 NMR chemical shifts from a single reference molecular geometry. Current ML methods for chemical property prediction utilize a single molecular geometry as model inputs. However, in real-world applications, a single molecule can adopt a variety of different geometries, as dictated by its Boltzmann distribution; the experimentally measured property is a weighted average of these values. Thus, the prediction results of current models fail to reproduce real-world experimental results accurately. For instance, the chemical shifts of carbon atoms in a molecule are dependent on molecular geometries. This dependence ties the accuracy of predictive ML models[1] to the quality of the starting geometry. We will develop an ML model that encodes the geometric variability of molecules, allowing for more accurate predictions that better reflect experimental results.
Description of REU Student Activities:
- Implement a high-throughput workflow to compute DFT energies for a dataset of molecular conformers (Paton)
- Database management and developing chemical descriptors using Python packages like pandas and cheminformatics packages like RDKit (Kim)
- Develop an Equivariant Neural Network (ENN) ML model to predict molecular properties (Kim)
Mentors: Assistant Professor Ronnie Banerjee and Professor Debbie C. Crans
Project Description:
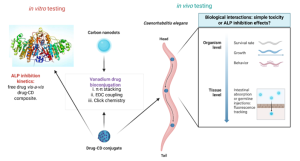
In the past, we have optimized the synthesis of water- and serum-soluble carbon nanodots (CDs) with tunable emissions and the capacity to support vanadium complexes through hydrophobic and π-π stacking interactions. These vanadium compounds are promising pharmaceuticals with anti-neoplastic and insulin-mimetic activity, with many superior candidates at various stages of development in the laboratory-to-pharmacy pipeline. The drug-CD conjugates represent a novel class of theragnostic materials combining diagnostic imaging with potential therapeutic applications in disease states such as cancer and diabetes. In a continuation of this work, we intend to examine alternative bioconjugation strategies such as click reactions and amide couplings for immobilization of the vanadium drugs onto the CDs. These conjugates are then to be compared to and contrasted with uncoordinated vanadium complexes in terms of their inhibition of the alkaline phosphatase (ALP) enzymes in vitro and extended eventually to simple organism models such as C elegans.
Description of REU Student Activities:
- Synthesis of the vanadium drugs and the CDs – Crans/Banerjee labs.
- Bioconjugation of the vanadium drugs to CDs – Banerjee lab.
- Characterization of the conjugates through one or more of the following techniques: UV-Vis-NIR spectroscopy, electron microscopy, dynamic light scattering, Raman spectroscopy, X-ray photoelectron spectroscopy, and fluorimetry – Banerjee lab and ARC.
- Release profile of the vanadium drugs from the conjugates – Crans lab.
- Enzyme kinetics for the inhibition of ALP by the drug-CD conjugates vis-à-vis uncoordinated vanadium complexes – Crans lab.
Mentors: Professors Jamie Neilson and Amy Prieto
Project Description:
To enable next-generation, beyond Li-ion batteries, we require new materials. For example, sodium is extremely abundant and inexpensive, and sodium ion batteries have the potential to replace lithium in myriad applications. However, we cannot simply replace Li with Na in existing materials used in litihium ion batteries. In this project, we are taking advantage of new synthetic solid-state chemistry to synthesize and test new materials for the development of sodium ion batteries.
Description of REU Student Activities:
- Synthesis of new ceramic materials using high-temperature furnaces.
- Characterization of materials using diffraction and electron microscopy.
- Construction of electrochemical cells
- Electrochemical testing of materials in battery half-cells.
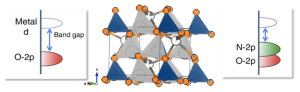
Mentors: Professors Jamie Neilson and Justin Sambur
Project Description:
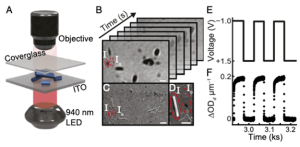
This project will use a new optical microscope in the Sambur Lab, QSCAT, to image single particles and measure ion concentrations in battery materials. QSCAT makes it possible to directly link structure and composition at the single-particle level, shedding light on why some particles act as ‘champions’ while others ‘fail’ in a working battery.
Description of REU Student Activities:
- The student will synthesize solid-state battery compounds in the Neilson Lab and carry out single-particle measurements in the Sambur Lab.
- Through this work, the student will develop expertise in materials synthesis, microscopy, spectroscopy, and X-ray diffraction.
- The student will also gain computational skills by analyzing experimental data using MATLAB.
Mentors: Professors Garret Miyake and Robert Paton
Project Description:
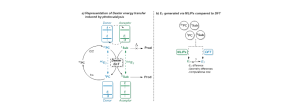
Dexter energy transfers (EnT), predominant processes encountered in photocatalysis, correspond to the formation of a triplet state photocatalyst (donor) which transfers energy to the ground state of the substrate (acceptor), resulting in a triplet substrate and recovering of the ground state photocatalyst (Figure a). The energy difference between the singlet ground state and the lowest triplet state (triplet energy or ET) of the photocatalyst and the substrate is a good descriptor to rationalize the rate of such reactions. While these triplet energies are routinely modeled and predicted by density functional theory (DFT), recent machine learning interatomic potentials (MLIPs) could considerably reduce the computational cost associated with these calculations, suggesting the opportunity for prediction and/or catalyst design. This project will focus on assessing the efficiency of MLIPs to predict triplet energy by comparing their accuracy with DFT results of previously published systems (Figure b). Those systems will be chosen so that an array of organic photocatalysts and substrates partners will be represented, in order to identify potential functional groups for which MLIPs are the most appropriate and others for which a different approach should be preferred.
Description of REU Student Activities:
- Short literature review on the subject
- Introduction to Linux environment
- Introduction to density functional theory and machine learning
- Development of a workflow
- Results analysis and presentation